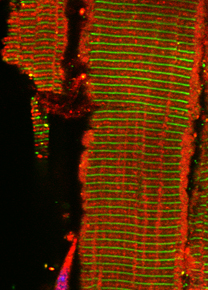
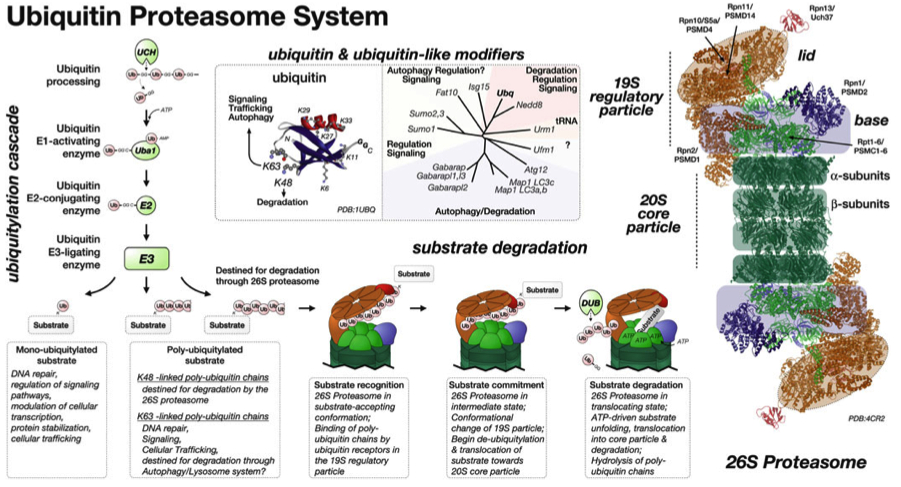
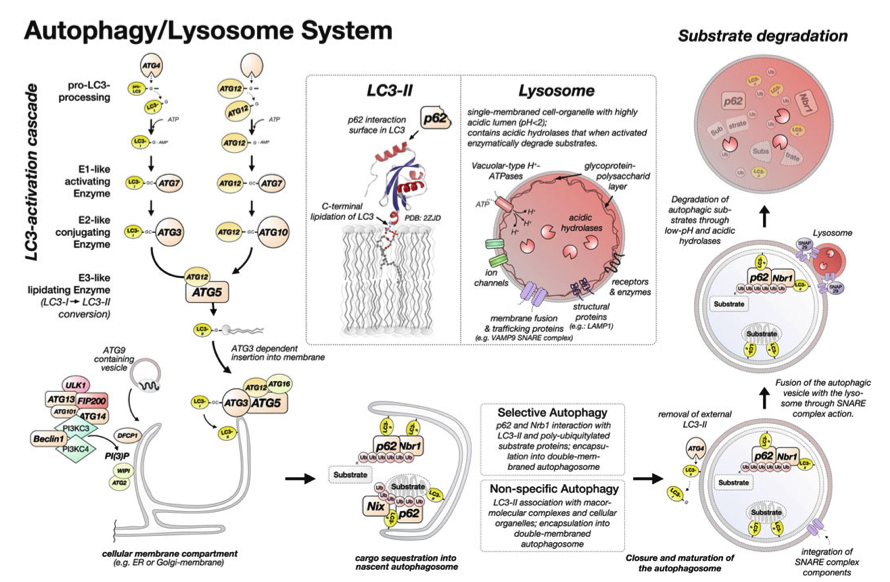
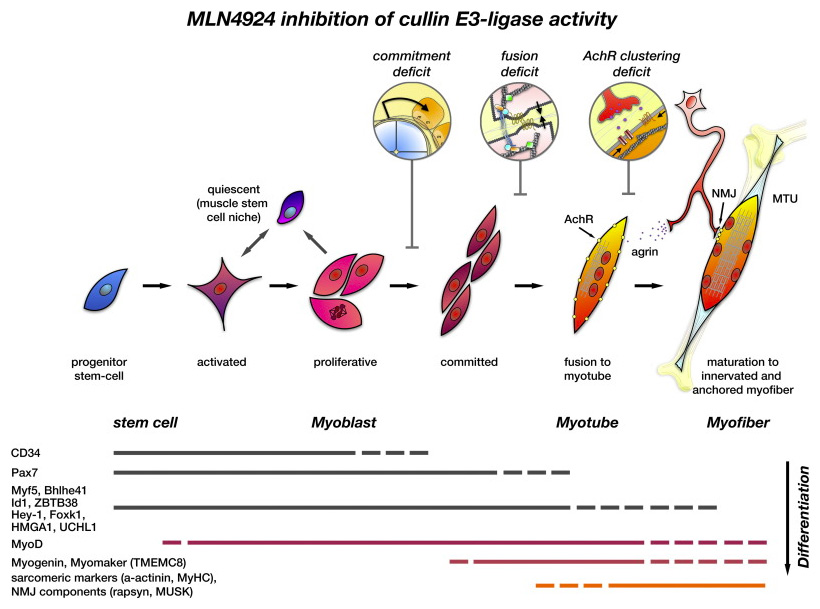
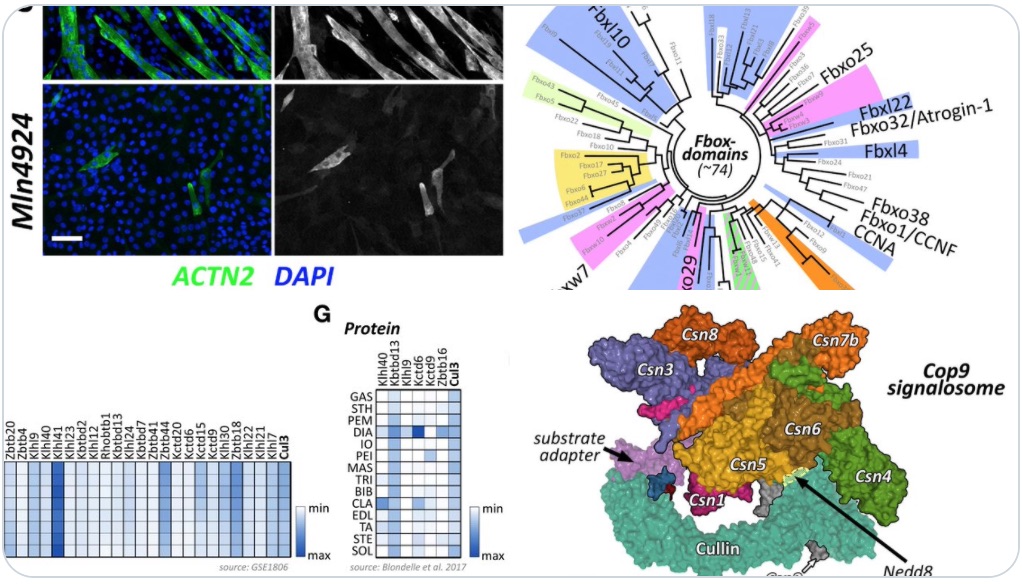
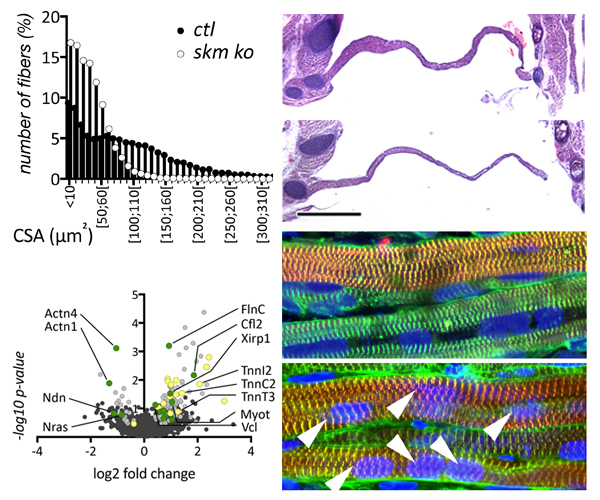
In our new manuscript, we outlined the role that Cullin3 E3-ligase function plays for the development of nemaline myopathy in a murine model of the disease. Using mice that are deficient of Cullin-3, we define a novel molecular pathomechanism that may be at play in patients suffering from mutations in KBTBD13. The manuscript can be found at JCI Insight.
Main objectives are to study:
- further characterize muscle specific functions for E3-ligases of the cullin family
- identify substrate-adaptor proteins for cullin-3
- investigate the role for cullin-3 for myopathy development
Related Publications
- The kinase domain of titin controls muscle gene expression and protein turnover. Lange S, Xiang F, Yakovenko A, Vihola A, Hackman P, Rostkova E, Kristensen J, Brandmeier B, Franzen G, Hedberg B, Gunnarsson LG, Hughes SM, Marchand S, Sejersen T, Richard I, Edstroem L, Ehler E, Udd B, Gautel M. Science. 2005 Jun 10;308(5728):1599-603. PMID: 15802564
- Obscurin and KCTD6 regulate cullin-dependent small ankyrin-1 (sAnk1.5) protein turnover. Lange S, Perera S, Teh P, Chen J. Mol Biol Cell. 2012 Jul;23(13):2490-504. PMID: 22573887
- Breaking down protein degradation mechanisms in cardiac muscle. Lyon RC, Lange S, Sheikh F. Trends Mol Med. 2013 Apr;19(4):239-49. PMID: 23453282
- Cardiac Cytoarchitecture: How to Maintain a Working Heart-Waste Disposal and Recycling in Cardiomyocytes. Jordan Blondelle & Stephan Lange. Book Chapter in Cardiac Cytoarchitecture. Edited by Elisabeth Ehler. Springer International Publishing Switzerland. 2015. ISBN: 978-3-319-15262-2 (Print) 978-3-319-15263-9 (Online). Online Version
- Cullin E3 ligase activity is required for myoblast differentiation. Jordan Blondelle, Paige Shapiro, Andrea A. Domenighetti, Stephan Lange. JMB. 2017. dx.doi.org/10.1016/j.jmb.2017.02.012
- Cullin-3 dependent deregulation of ACTN1 represents a new pathogenic mechanism in nemaline myopathy. Blondelle J, Tallapaka K, Seto JT, Ghassemian M, Clark M, Laitila JM, Bournazos A, Singer JD, Lange S. JCI Insight. 2019 Apr 16. doi.org/10.1172/jci.insight.125665
- The Role of Cullin-RING Ligases in Striated Muscle Development, Function, and Disease. Blondelle J, Biju A, Lange S. IJMS. 2020, 21(21), 7936. doi.org/10.3390/ijms21217936
Future Directions
We are currently investigating whether E3-ligases of the cullin protein family play a greater role for muscle specific protein turnover than previously anticipated. Of particular interest is the possible role that cullin-3 plays for the development of skeletal and cardiac muscles as well as motor neurons. We also study pathophysiological roles of cullin-3 for the etiology of neuromuscular diseases and cardomyopathies.
Collaborators on this project
- Dr. Jeffrey Singer at Portland State University, OR
- Dr. Jane Seto at Murdoch Children's Hospital, Australia
- Drs. Careina Wallgren-Pettersson & Katarina Pelin at the University of Helsinki, Finland
Keywords: degradation, cullin, E3-ligase, ubiquitin.
|